Non-native fish can cause major damage to New Zealand lakes, so detecting them early is crucial for protecting local ecosystems. Traditional netting or trapping methods often miss new or low-abundance populations, prompting growing interest in environmental DNA (eDNA) — a sensitive method that detects species by analysing fragments of genetic material left behind in the water or sediment.
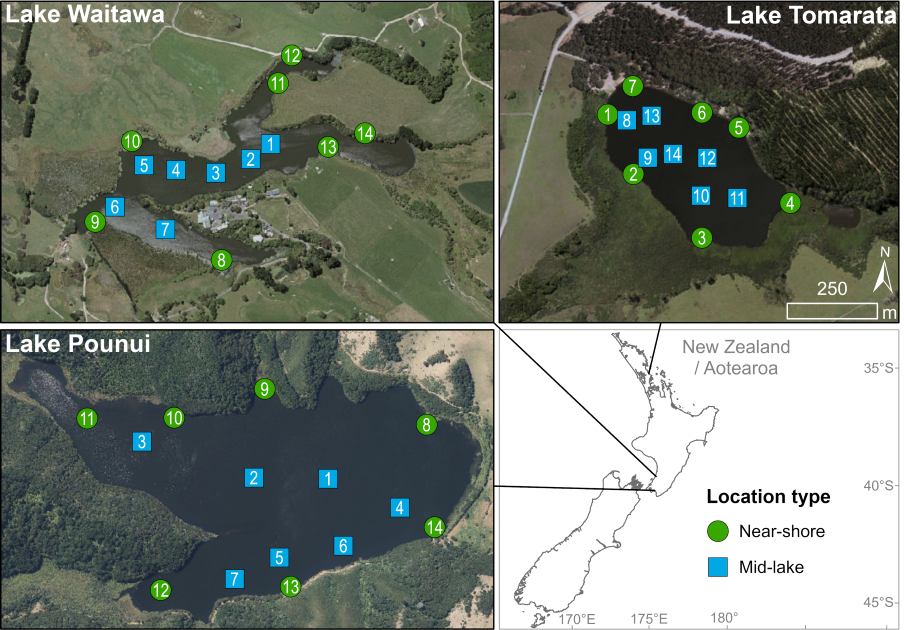
This study tested how best to use eDNA to find two invasive fish species — European perch and rudd — in three small shallow lakes. Researchers compared water samples and sediment samples collected from both the lake edges and mid-lake locations. They then used advanced digital PCR methods and probabilistic modelling to determine how many samples are needed for reliable detection.
The findings show that sample type and sampling effort matter. To confidently detect fish DNA in sediment, monitoring programs should include at least six sites with five replicates each. For water samples, detection requires even more effort — twelve sites with eight replicates each. These results emphasise that effective eDNA surveillance needs lake-specific sampling strategies, tailored to habitat features and lake size.
Read the full paper here.