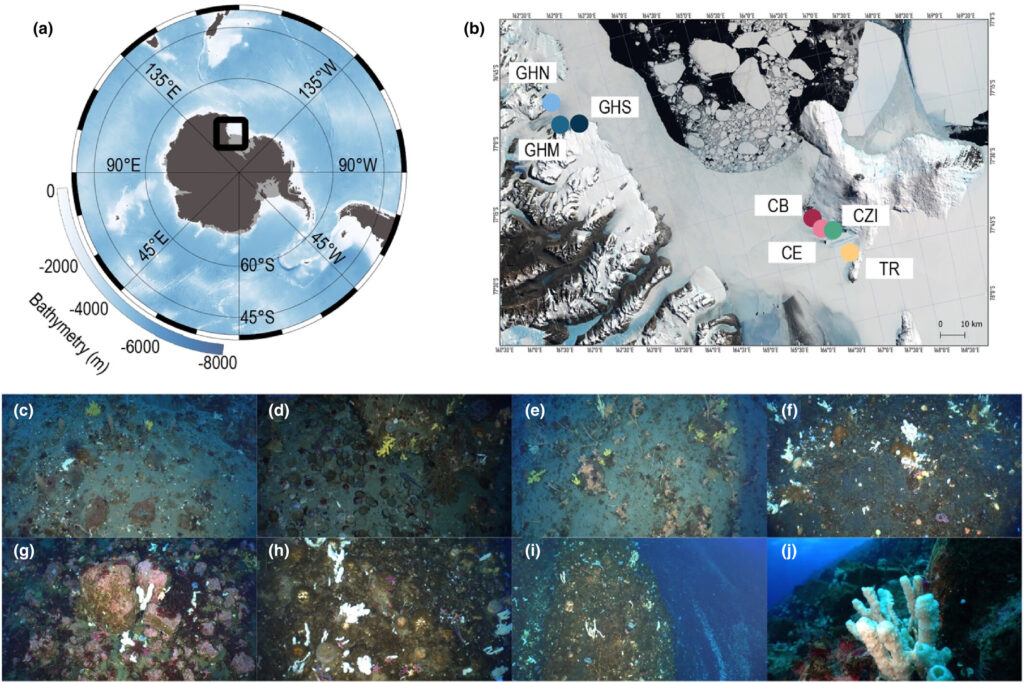
The Ross Sea in Antarctica is one of the planet’s most pristine marine environments, but growing human activity is increasing the need for effective biodiversity monitoring. Traditional surveys in such remote, icy waters are difficult and resource-intensive. This study explores whether environmental DNA (eDNA) collected from both seawater and marine sponges can provide a more practical way to monitor life in this unique region.
Researchers sampled seven coastal sites, using metabarcoding to analyse DNA traces from both substrates. Across all samples, they detected an impressive 1,450 eukaryotic taxa spanning 30 phyla, revealing the extraordinary diversity of life in the Ross Sea. However, the results showed that sponges and seawater capture different parts of the ecosystem, with only partial overlap between the two. Sponge eDNA often reflects organisms that interact closely with the sponge or settle on its surface, while seawater provides a broader snapshot of the surrounding community.
The study also highlighted an ongoing challenge in polar biodiversity research: limited reference DNA databases. Only 9% of detected taxa could be confidently identified to species level, and 40% could not be classified at all. Despite this, clear differences in biodiversity between sites matched existing knowledge of species distributions, demonstrating the power of eDNA to detect real ecological patterns.
Overall, the study shows that eDNA offers a promising, low-impact way to monitor remote Antarctic ecosystems. To get the most complete picture, however, multiple sample types — such as seawater and sponge tissues — should be combined. The work also underscores the need for more comprehensive Antarctic reference genomes to unlock the full potential of eDNA monitoring.
Read the full article here.