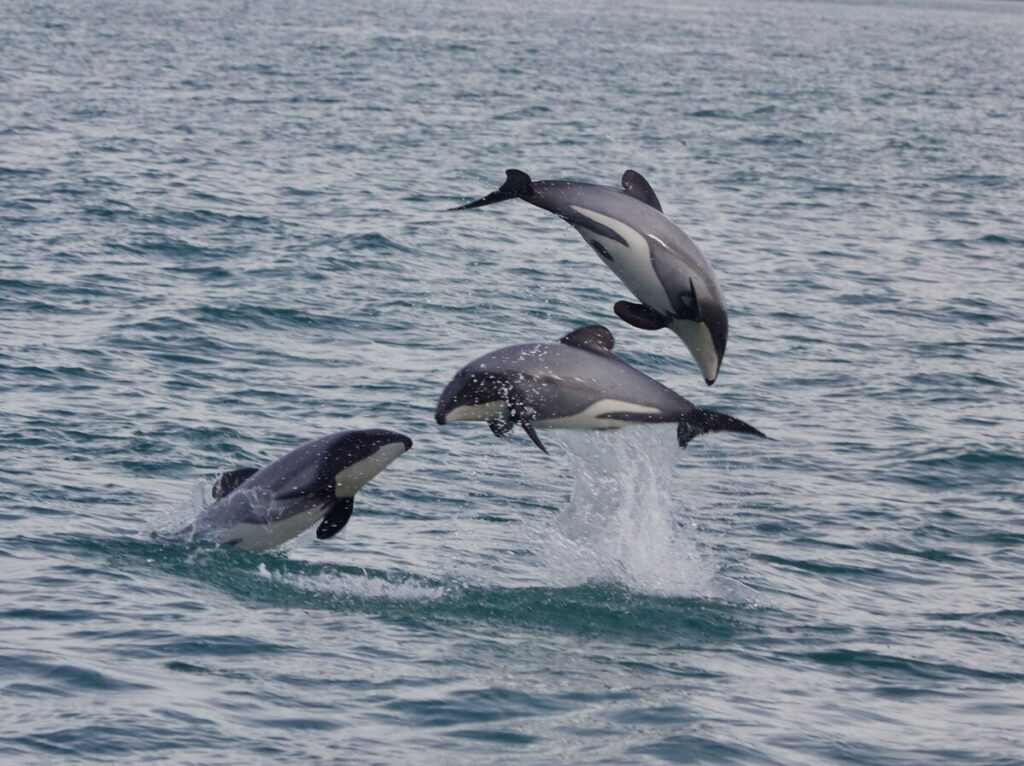
Hector’s dolphin, an endemic and culturally significant (taonga) species in Aotearoa New Zealand, is known to consist of several regional subpopulations, but traditional genetic studies rely on tissue samples that are difficult and invasive to collect. This study demonstrates how environmental DNA (eDNA) can help fill that gap by detecting population-level genetic variation directly from seawater.
Researchers collected 85 water samples across three East Coast locations during the 2022/23 summer: Banks Peninsula, Timaru, and Dunedin. Using a mitochondrial DNA marker commonly applied in dolphin genetics, they successfully identified seven distinct haplotypes (genetic lineages) from the eDNA. Importantly, the team developed a simple method to remove PCR and sequencing errors, enabling true haplotypes to be distinguished from noise even when DNA from multiple individuals is mixed together.
The results closely matched decades of tissue-based research for Banks Peninsula and Timaru, confirming that eDNA can reliably reproduce known population patterns. The Dunedin samples, however, told a different story: their haplotypes more closely resembled dolphins from the South Coast than those from the East Coast South Island group. This finding suggests that the boundaries of the East Coast subpopulation may need to be reconsidered, which has direct implications for management and protection strategies.
Read the full paper here.
Photo credit: Hector’sDolphinsCloudyBay 21Feb2012 AnjanetteBaker.tif. (2025, May 31). Wikimedia Commons. Retrieved December 12, 2025, from https://commons.wikimedia.org/w/index.php?title=File:Hector%27sDolphinsCloudyBay_21Feb2012_AnjanetteBaker.tif&oldid=1038629124.