eDNA is revolutionising biodiversity monitoring, helping detect endangered species in remote habitats. But it’s also proving invaluable for taxonomists, offering genetic insights that can reshape the tree of life itself.
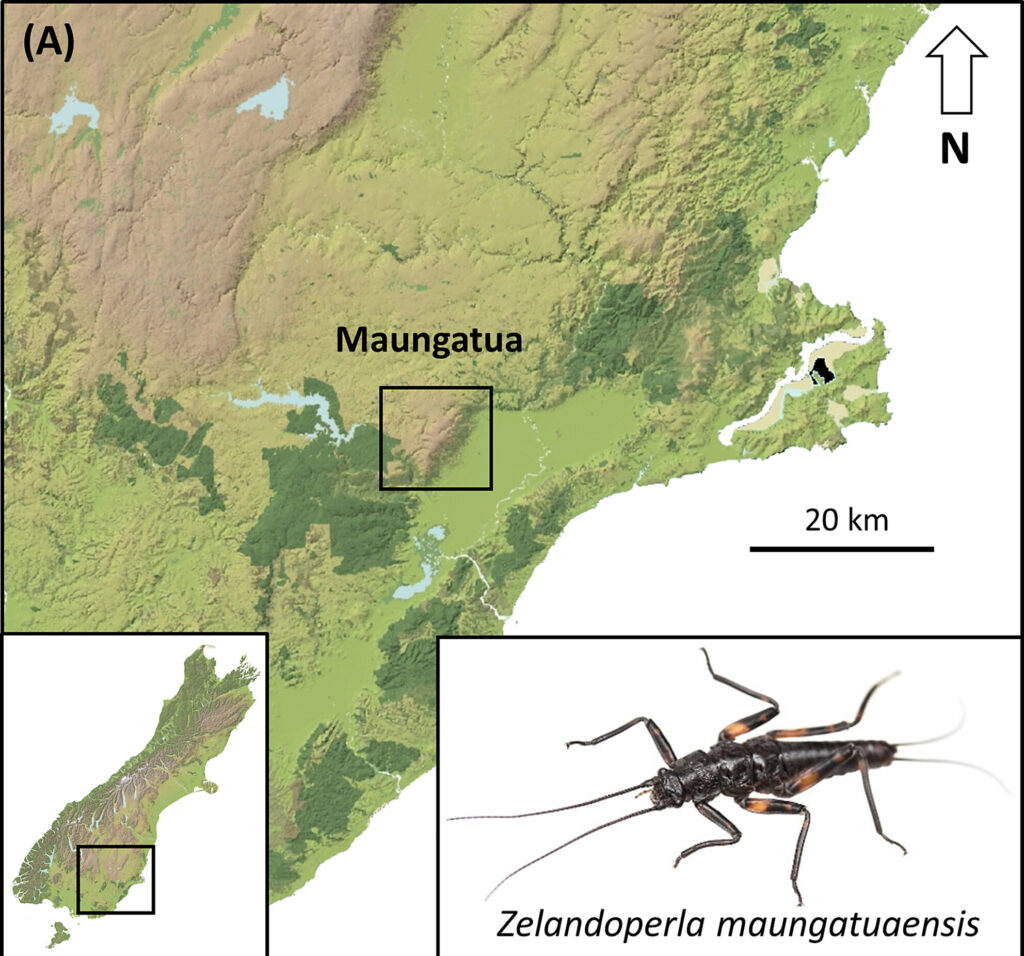
A recent study led by Graham McCulloch, and with Steve Pohe, Tom Drinan and Jon Waters, aimed to assess the geographic distribution of the Maungatua stonefly, Zelandoperla maungatuaensis. This rare species had been previously found in only a handful of streams in the remote mountain ranges of southern New Zealand. Locating these populations is crucial for protecting the species but accessing remote upland streams can be difficult and these small invertebrates can be tricky to sample and identify.
Using targeted eDNA metabarcoding, the team not only confirmed the presence of the few known stonefly populations but also detected two previously undiscovered populations in nearby streams. What’s more, the eDNA sequences from these new sites showed subtle differences from known reference data. Follow-up fieldwork confirmed the presence of a previously unknown, genetically distinct subclade.
This case highlights the fast-evolving synergy between eDNA and taxonomy. While robust eDNA monitoring relies on accurate, up-to-date reference data, eDNA itself can play a vital role in closing the gaps in our taxonomic knowledge. Every new identification improves the data available for future monitoring efforts, enabling conservationists to better understand and protect isolated species, like the Maungatua stonefly, in all their genetic diversity.
Want to learn more? Read the full paper here.