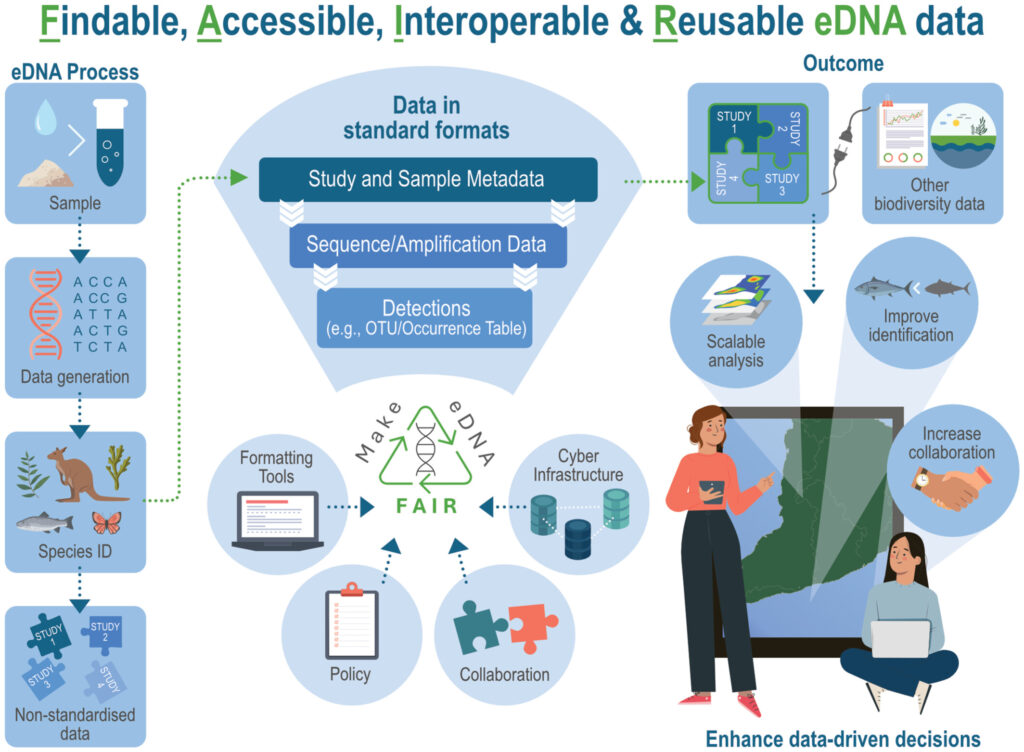
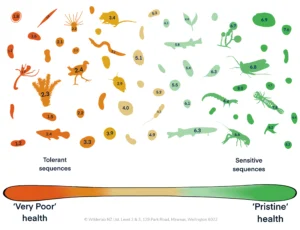
Environmental DNA (eDNA) has transformed biodiversity monitoring by making it possible to detect species and assess whole communities from small water, soil, or sediment samples. However, the enormous value of these datasets is often limited by inconsistent formatting, missing metadata, and restricted access. When eDNA data are difficult to find, compare, or reuse, their long-term scientific and management impact is reduced. To address this challenge, this paper introduces a comprehensive framework to make eDNA data FAIR: Findable, Accessible, Interoperable, and Reusable.
The authors present the FAIR eDNA (FAIRe) Metadata Checklist, which adapts and expands existing standards such as Darwin Core (DwC) and MIxS to cover the specific needs of eDNA workflows. The checklist links all stages of eDNA data—from raw sequence files and qPCR values to processed ASV/OTU tables and species detections—to fully described metadata. The accompanying formatting guidelines, templates, and example datasets provide a practical pathway for researchers, managers, and data stewards to adopt FAIR practices immediately.
By standardizing how eDNA data are described and shared, the approach aims to make biodiversity information more accessible, comparable across studies, and ready for large-scale analyses. The paper also outlines a long-term plan to integrate these guidelines into global biodiversity data standards and community practices. Widespread adoption of FAIR principles will support more robust biodiversity assessments, improve species distribution mapping, and enable broader ecological insights from the rapidly growing body of eDNA data.
Read the full article here.