Wetlands are culturally and ecologically important ecosystems, but they are also among the world’s most threatened. While eDNA biomonitoring is well established in rivers, lakes, and marine systems, wetlands pose unique challenges because high sediment loads clog filters and make water collection difficult. This study explored how eDNA behaves in a New Zealand wetland and tested practical sampling approaches to help guide future monitoring in these complex environments.
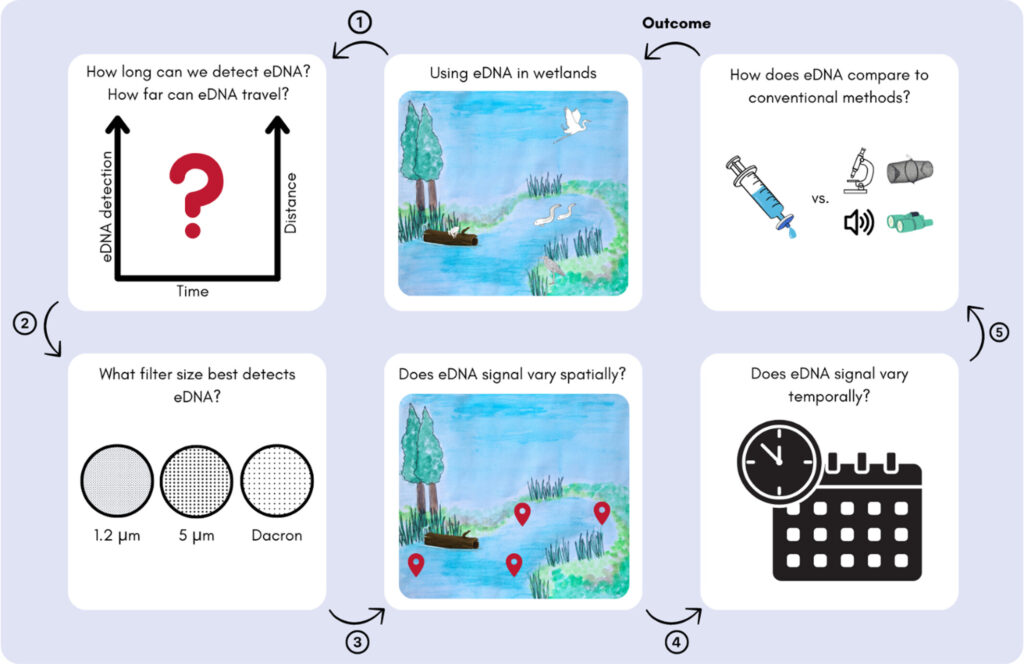
Researchers sampled four sites within the Opuatia Wetland during three springtime surveys, using conventional taxonomic methods alongside multiple eDNA filtration approaches (1.2 µm, 5 µm, and semi-quantitative dacron filters). They also conducted a controlled experiment releasing trace DNA from kea to track how far and how long it could be detected.
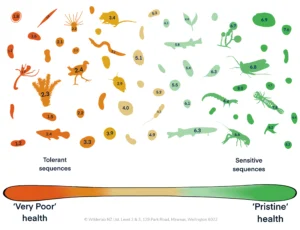
The results showed clear spatial and temporal shifts in biodiversity, and notable differences in the species detected using each filter type. eDNA complemented, rather than replaced, conventional surveys, with each method detecting species the other missed. Foreign DNA remained detectable up to 10 metres from the release point for as long as one week, offering insight into eDNA persistence and movement in wetland environments.
Overall, this research provides important considerations for designing wetland eDNA programs. It highlights the need to account for sediment-rich water, variable DNA persistence, filter performance, and the value of combining eDNA with traditional survey methods.
Read the full paper here.